Density-Matrix Embbeding Theory (DMET)
Vayesta can be used for DMET calculations of molecules, solids, and lattice models. In this section, we give two simple examples: the calculations of a \(\textrm{H}_6\) molecule and a 1D Hubbard model.
Simple Molecule
A simple DMET calculation of a \(\textrm{H}_6\) ring can be performed as follows (the example can also be found in examples/dmet/01-simple-dmet.py):
1import pyscf.gto
2import pyscf.scf
3import pyscf.fci
4import vayesta
5import vayesta.dmet
6from vayesta.misc.molecules import ring
7
8
9# H6 ring
10mol = pyscf.gto.Mole()
11mol.atom = ring(atom="H", natom=6, bond_length=2.0)
12mol.basis = "sto-6g"
13mol.output = "pyscf.out"
14mol.build()
15
16# Hartree-Fock
17mf = pyscf.scf.RHF(mol)
18mf.kernel()
19
20# Reference FCI
21fci = pyscf.fci.FCI(mf)
22fci.kernel()
23
24# One-shot DMET
25dmet = vayesta.dmet.DMET(mf, solver="FCI", maxiter=1)
26with dmet.sao_fragmentation() as f:
27 f.add_atomic_fragment([0, 1])
28 f.add_atomic_fragment([2, 3])
29 f.add_atomic_fragment([4, 5])
30dmet.kernel()
31
32# Self-consistent DMET
33dmet_sc = vayesta.dmet.DMET(mf, solver="FCI")
34with dmet_sc.sao_fragmentation() as f:
35 f.add_atomic_fragment([0, 1])
36 f.add_atomic_fragment([2, 3])
37 f.add_atomic_fragment([4, 5])
38dmet_sc.kernel()
39
40print("Energies")
41print("========")
42print(" HF: %+16.8f Ha" % mf.e_tot)
43print(" FCI: %+16.8f Ha" % fci.e_tot)
44print(" DMET(1 iteration): %+16.8f Ha (error= %.1f mHa)" % (dmet.e_tot, 1000 * (dmet.e_tot - fci.e_tot)))
45print(" DMET(self-consistent): %+16.8f Ha (error= %.1f mHa)" % (dmet_sc.e_tot, 1000 * (dmet_sc.e_tot - fci.e_tot)))
In lines 9–17 the \(\textrm{H}_6\) molecule is set up and a restricted Hartree–Fock (RHF) calculation is performed using PySCF. We also perform an full system full configuration interaction (FCI) calculation (lines 20–21), to obtain a reference value for the DMET results.
In line 24, a one-shot DMET calculation is instantiated with the Hartree–Fock object as first argument.
Additionally, the keyword argument solver='FCI' is used to select FCI
as the default solver and maxiter=1 is used to skip the DMET self-consistency cycle.
In lines 25–28 the fragments for the calculation are defined.
This is done with help of the context manager dmet.sao_fragmentation(), which specifies that the
fragments added in the body of the context manager will refer to symmetrically orthogonalized atomic orbitals (SAOs).
The method f.add_atomic_fragment(atoms) adds a fragment comprised of
all orbitals corresponding to the atom atoms to the calculation.
In this case, we split the system into three fragments, containing two neighboring hydrogen atoms each.
Note
For more information on the fragmentation and the add_atomic_fragment method,
see section Defining Fragments.
Finally, the two embedding problems are solved by calling dmet.kernel()
and the resulting total energy is stored in dmet.e_tot.
In lines 32–37 these steps are repeated for a self-consistent DMET calculation, in which case only
the keyword argument maxiter=1 has to be dropped.
Note
The fragments in the calculation above are symmetry-related by a rotation of 120° or 240° around the z-axis. This symmetry can be added to the embedding class, such that only a single fragment needs to be solved:
# One-shot DMET
dmet = vayesta.dmet.DMET(mf, solver="FCI", maxiter=1)
with dmet.sao_fragmentation() as f:
with f.rotational_symmetry(3, axis=[0, 0, 1]):
f.add_atomic_fragment([0, 1])
dmet.kernel()
see also example/dmet/02-rotations-symmetry.py
Note
The self-consistent optimization of a correlation potential in DMET can be ill-defined and will sometimes fail. In this case, changing the tolerance or starting point can sometimes help. The underlying origin of these difficulties are described in Ref. [1].
Our implementation of self-consistent DMET follows that of Ref. [2] and requires the Python package cxvpy
for use.
Density-Matrices
In DMET expectation values are defined in terms of democratically partitioned one- and two-body density-matices.
These can be obtained by calling dmet.make_rdm1() and dmet.make_rdm2() as shown in example
examples/dmet/03-density-matrices.py:
1import numpy as np
2import pyscf.gto
3import pyscf.scf
4import vayesta
5import vayesta.dmet
6from vayesta.misc.molecules import ring
7
8# H6 ring
9mol = pyscf.gto.Mole()
10mol.atom = ring(atom="H", natom=6, bond_length=2.0)
11mol.basis = "sto-6g"
12mol.output = "pyscf.out"
13mol.build()
14
15# Hartree-Fock
16mf = pyscf.scf.RHF(mol)
17mf.kernel()
18
19# One-shot DMET
20dmet = vayesta.dmet.DMET(mf, solver="FCI", maxiter=1)
21with dmet.sao_fragmentation() as f:
22 f.add_atomic_fragment([0, 1])
23 f.add_atomic_fragment([2, 3])
24 f.add_atomic_fragment([4, 5])
25dmet.kernel()
26
27# Calculate energy from democratically-partitioned density matrices:
28dm1 = dmet.make_rdm1(ao_basis=True)
29dm2 = dmet.make_rdm2(ao_basis=True)
30# One and two-electron integrals in AO basis:
31h1e = dmet.get_hcore()
32eris = dmet.get_eris_array(np.eye(mol.nao))
33e_dmet = dmet.e_nuc + np.einsum("ij,ij->", h1e, dm1) + np.einsum("ijkl,ijkl->", eris, dm2) / 2
34
35print("Energies")
36print("========")
37print(" HF: %+16.8f Ha" % mf.e_tot)
38print(" DMET: %+16.8f Ha" % dmet.e_tot)
39print(" DMET (from DMs): %+16.8f Ha" % e_dmet)
Hubbard Model in 1D
In order to simulate lattice model systems, such as the Hubbard model, Vayesta provides the classes
Hubbard1D, Hubbard2D and LatticeMF in the lattmod package.
In the following example the half-filled, ten-site Hubbard chain is calculated with \(U = 6t\) (where \(t\) is the hopping parameter, which is 1 by default):
1import numpy as np
2import vayesta
3import vayesta.dmet
4import vayesta.lattmod
5
6
7nsite = 10
8nimp = 2
9hubbard_u = 6.0
10boundary = "PBC"
11nelectron = nsite
12mol = vayesta.lattmod.Hubbard1D(nsite, nelectron=nelectron, hubbard_u=hubbard_u, boundary=boundary)
13mf = vayesta.lattmod.LatticeMF(mol)
14mf.kernel()
15
16# Calculate each 2-sites fragment:
17dmet = vayesta.dmet.DMET(mf, solver="FCI")
18with dmet.site_fragmentation() as f:
19 for site in range(0, nsite, nimp):
20 f.add_atomic_fragment(list(range(site, site + nimp)))
21dmet.kernel()
22
23print(dmet.fragments)
24# Calculate a single fragment and use translational symmetry:
25dmet_sym = vayesta.dmet.DMET(mf, solver="FCI")
26# Specify the number of translational copies in direction of the three lattice vectors by passing a list with three
27# integers, [n0, n1, n2]. 1D or 2D systems have their periodic dimension along the first one or two axes.
28nimages = [nsite // nimp, 1, 1]
29dmet_sym.symmetry.set_translations(nimages)
30# Add only a single 2-sites fragment:
31with dmet_sym.site_fragmentation() as f:
32 f.add_atomic_fragment(list(range(nimp)))
33dmet_sym.kernel()
34
35print("Difference in converged solutions:")
36print(" |d(E_tot)|= %.5e" % abs(dmet.e_tot - dmet_sym.e_tot))
37print(" |d(V_corr)|= %.5e" % np.linalg.norm(dmet.vcorr - dmet_sym.vcorr))
For lattice model systems, fragments for quantum embedding calculations are usually defined in terms of lattice sites.
For this purpose, the embedding class has the fragmentation context manager dmet.site_fragmentation().
Within the body of this context manager, fragments can be added as before with the method
add_atomic_fragment—for the purpose fo defining fragments, the sites are considered as atoms.
In lines 19–20 of this example, the lattice is divided into two-site fragments, as depicted in Fig. 2.
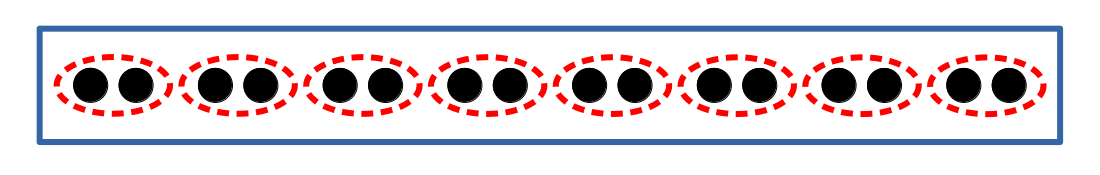
Fig. 2 Schematic depiction of the 1-D Hubbard model with two-sites fragments.
Just as for the \(\mathrm{H}_6\) ring molecule in the example above, this lattice model system has an inherent (in this case translational) symmetry between the fragments, which can be exploited. This is done in the second DMET calculation in lines 24–32, where the translational symmetry is specified in the following lines:
# integers, [n0, n1, n2]. 1D or 2D systems have their periodic dimension along the first one or two axes.
nimages = [nsite // nimp, 1, 1]
The three integers in nimages specify the number of symmetry related copies (including the original)
along each lattice vector.
For a 1D system, the first lattice vector corresponds to the periodic dimension and is thus the only dimension
along which there are more than one copies.